In our popular culture we have a complicated relationship with mutants. For every Teenage Mutant Turtle and Wolverine, there is a Magneto, a Jason Vorhees or a Thing. The hillbillies from Deliverance probably qualify as mutants but I’m not sure whether Godzilla is a good guy or a bad guy? Are zombies mutants? Regardless, it seems that everyone agrees that messing with your DNA is a risky proposition. Sure, you might get some superpowers, but you can also end up a horrifyingly deformed monstrosity, someone who isn’t kind to visiting city slickers or a fly, if your Jeff Goldblum.

Don’t mess with your DNA seems like good advice. But, as the boring scientist in the room, I should point out that mutants are not that rare. In fact, you can you make a strong case that every last one of us is a mutant. Every gene with multiple versions, the versions that give us blue or brown eyes for example, is the result of a mutation sometime in the past. Not only that, our DNA is full of mistakes that didn’t exist in our parents1. Every single one of us has around 60-70 new mutations that our parents don’t possess, and that are unique to us and potentially our children.
So, mutations aren’t rare. They happen all the time. In fact, they are the driving force behind genetic diversity. The reservoir of traits in our population that could come in handy if we, as a species, encounter something new. Something like climate change, a new disease or a zombie apocalypse. Without genetic diversity, for example, there wouldn’t be that one person who is resistant to the zombie infection. You know, the one who must be protected at all costs and taken to the CDC so scientists can make a zombie vaccine. Genetic diversity, and the little differences in DNA that defines it, makes every human respond a little differently to the world around us and those differences could make all the difference for our survival as a species.
If we are looking for an area where we can have a look at some practical examples of human genetic diversity we need look no further than our dinner plates. Zombie apocalypses are rare, but we eat every day and we have a lot of biological machinery that digests and metabolises that food. All of that machinery, ultimately, relies on the information encoded in our genome for its functioning, which means there will be individual variations in our metabolisms due to genetic diversity.
We also have food preferences. Different foods that different individuals prefer to eat. While we tend to think of these preferences as the result of learning and upbringing, there is more and more evidence that genetics also plays an important role. That is, maybe we love mum’s cooking because she also gave us many of our genes, including some of the gene variants that shaped her own food preferences.
Before we look at some of these food-based genetic variants we need to have some understanding of how mutations affect us and how they occur. Genetics is a big, complicated subject but a simple way of thinking about it is that our genome encodes information that allows us to produce proteins. Proteins are the molecules that actually interact with the world. They are structural, they mediate chemical reactions and they serve as receptors on cells that kick off cellular reactions to the external environment. Mutations in our genome can alter the ways that these proteins function and thus the way our bodies are put together and the way that we react to different stimuli.
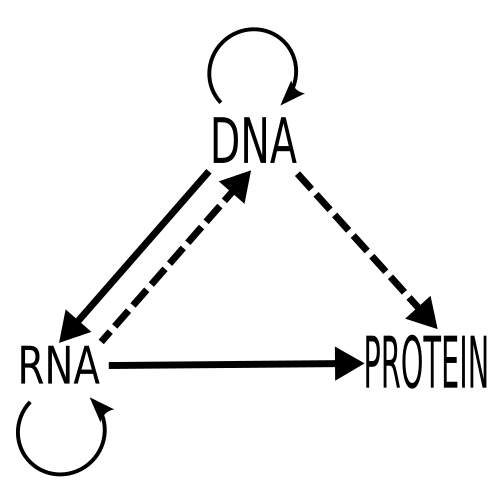
Almost all mutations occur when a cell is splitting to produce two daughter cells. To produce two cells the genome of the parent cell needs to be replicated so that you end up with one complete genome for each daughter cell. There is a whole cellular machinery that replicates the genome, but, like a medieval monk copying out manuscripts, it can make mistakes. One of the mistakes it can make is messing up the code in one spot2. Like the monk making a spelling mistake in a manuscript they are copying. These types of mutations are called single nucleotide polymorphisms (SNPs) and they are one of the most common mutations we find in the a human genome3.
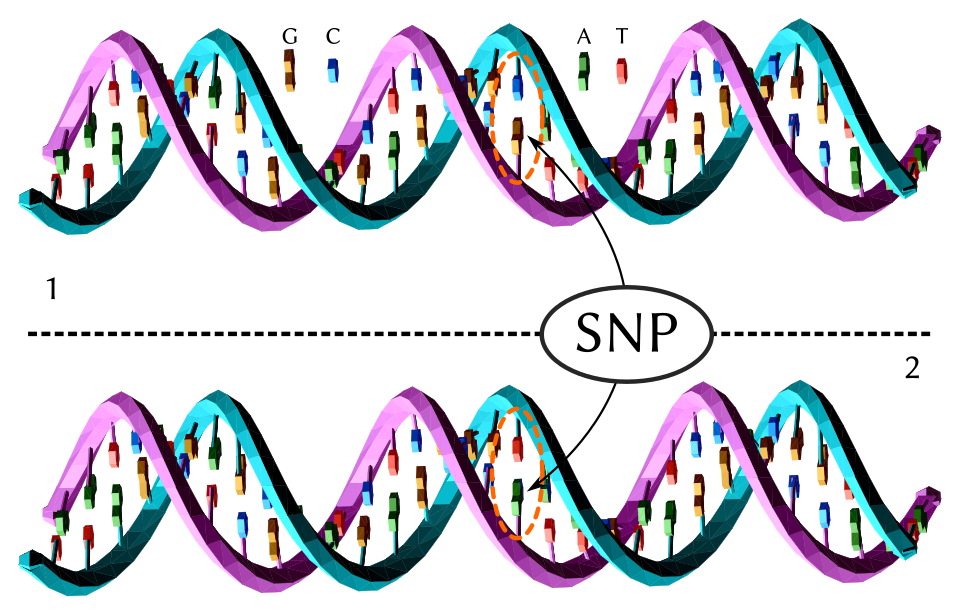
Probably one of the better known food-related gene variants is the “soapy” coriander4 mutation. We have the soapy coriander variant because sometime in the past someone experienced a SNP in a gene called OR6A2 and that mutation was passed onto that person’s descendants. Now, depending on the ethnic group, between 3-21% of the population can’t eat coriander because, to them, it tastes like soap, dirt or insects. A clue to why this happens is in the name of the gene. The “OR” in OR6A2 stands for “olfactory receptor” because this gene encodes one of the protein receptors that lines the cells of our nasal cavity and interacts with airborne molecules. When a molecule interacts with an olfactory receptor it triggers a nerve impulse that flows to the brain and is interpreted as an odour.
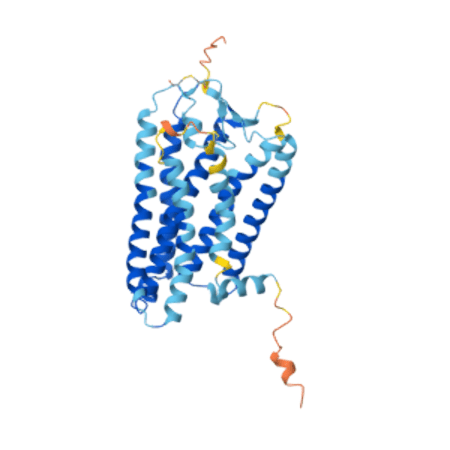
The OR6A2 receptor interacts with some types of a class of chemical called aldehydes. Aldehydes are common molecules in plants, and they stimulate a wide variety of aromas in humans, but some of them possess a “soapy” aroma and, you can probably see where this is going, they are present in small amounts in coriander. The change in the protein sequence caused by the SNP makes these aldehydes bind to the mutated receptor much more tightly than the “normal” OR6A2 receptor, stimulating more nervous signals to the brain. Most of us can’t detect these aldehydes in the small quantities you find in coriander but those with the “soapy” variant of the OR6A2 gene can. Essentially the SNP turns the normal OR6A2 into a “super detector”. I’ve written about the important role that smell has in perceiving flavour and the end result for those with the super-detecting OR6A2 is that their coriander tastes soapy5.
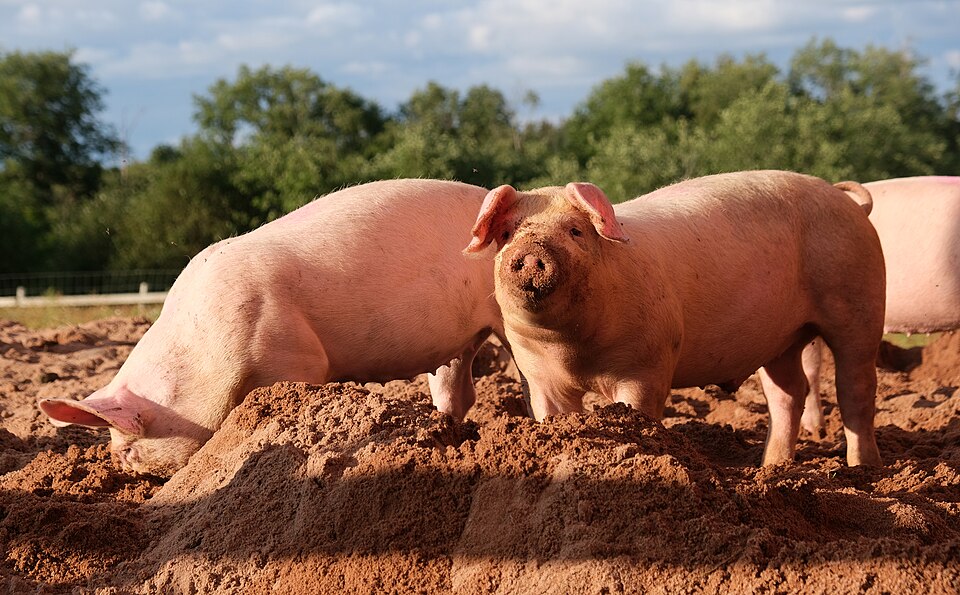
We have about 400 olfactory receptor genes so it seems reasonable to expect that variation in the 399 other OR genes might cause some peculiarities in peoples flavour response. And, sure enough, we find exactly that. One example is detecting boar taint, the smell, variously described as urine or faecal-like, that is associated with meat from uncastrated male pigs. The odour is primarily caused by andanole, a steroid, and skatole, another metabolic product, that accumulate in fatty tissue, producing the odour when the fat is heated.
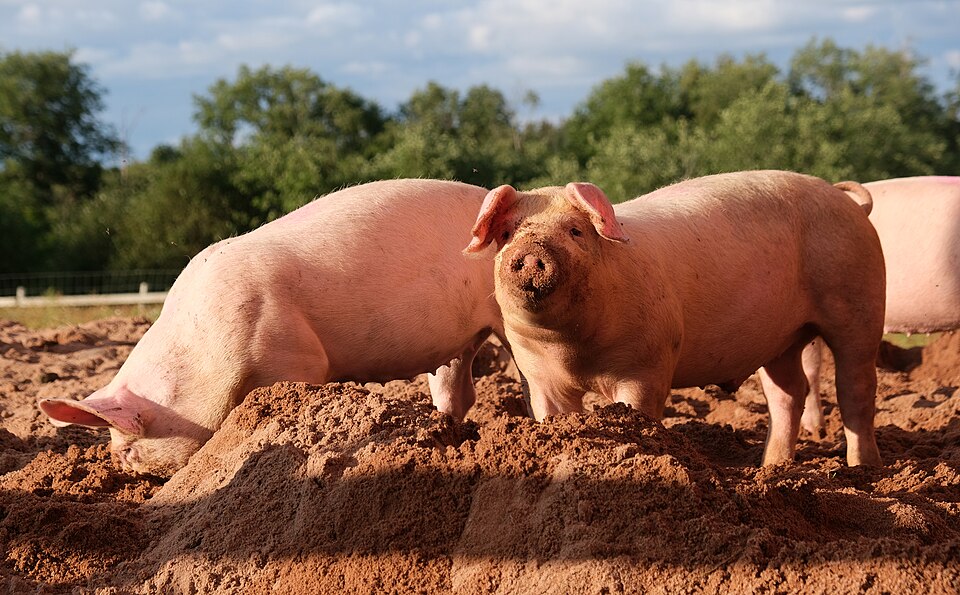

According to which source you look at, about 70% of the population are unable to detect the boar taint smell because of a variant of the OR7D4 gene. This variant has two SNPs and if you end up with one or two copies of this variant in your genome you are much less sensitive to boar taint. Unlike the soap coriander mutation, in this case the SNPs are reducing the effectiveness of the receptor rather than enhancing it, something to be thankful for if you ask me.
A SNP in the OR5A1 gene determines whether you perceive beta-ionone as a floral or harsh vinegary odour. Beta-ionone is found in wine but is also used in perfumes, which could cause problems if your date has this variant. At least three SNPs found in the OR2J3 gene can abolish the ability to smell cis-3-hexen-1-ol, which has the odour of freshly cut grass, and is found in tea, beer, wine and various fruits and vegetables. A single polymorphism in OR51B5 determines how easily you can smell cinnamon and other research has suggested a genetic basis for sensitivity to the odour of molecules found in blue cheese, teas and cocoa, red wine and a wide range of plant foods.
It’s not all olfactory genes though. As you’d expect variations in our taste receptors can also influence how intense a flavour we perceive in a given food. As taste receptors tend to stimulate a small set of “tastes” in reaction to many different types of molecules, bitter and sweet for example, variations can affect how much of certain types of food we will consume. This can have some important consequences for our health.
For example, variants in a receptor called CD36 that acts as a “fat” taste sensor in our taste buds has been linked to greater or lesser amounts of fat consumption. In mice, removing the CD36 gene removes the preference for fatty foods and in humans specific variants of CD36 have been associated with obesity (see here and here for the science). Because CD36 is down regulated when fatty foods are eaten, there is likely to be some type of feedback mechanism operating so the intensity of someone’s CD36 receptor could affect both the intensity of flavour as well as, potentially, short-circuiting the mechanisms meant to limit the amount of fat we want to eat.
Likewise, both our sweet taste buds and bitter taste receptors can vary significantly in the intensity of their response. This means that different people experience higher, or lower, levels of sweetness or bitterness from different foods. Variation in our sweetness receptors, and the intensity of our reaction to sugar, has been linked to differences in sugar consumption and the development of problems like obesity and diabetes (see here and here).
Alternatively, some people are bitterness super-tasters. In particular, variation in the TAS2R38 bitter receptor mediates sensitivity to bitter elements in our food. Those with two copies of a sensitive variant are extremely sensitive to bitterness and will generally avoid coffee, alcohol and, surprisingly, cigarettes (see here, here and here). They also avoid some more nutritional but bitter foods like Brussels sprouts, kale and the like, a less beneficial outcome.
If all this isn’t enough variation for us to consider. A recent large-scale study of food preferences used 161 625 participants to find 250 genes likely to be important in the formation of food preferences. Surprisingly, only 12 of the 250 genes identified were olfactory or taste receptors. Perhaps less surprisingly, the genes were predominantly associated with brain tissue. Suggesting that food preferences, if I can be forgiven for simplifying the issue, is mostly in our brain.
No single study is the final word on any subject, but this along with many other studies is starting to suggest that there is enormous variation in the way that we experience our food. Given the genetic variation that exists around flavour we really can’t be sure that what we are tasting is the same as what those around us are tasting. Which makes for some spirited discussion around food but also raises the question whether things like expensive wine can actually be discriminated by many who drink it?
It’s not just flavour though. Genetic variation can also affect the way that we digest and metabolise our food. I’ve already covered lactose intolerance where most of the world’s population can’t digest milk as adults. Also with a genetic basis is the way that we metabolise alcohol. Sometimes called the alcohol flush reaction, some people, particularly those of Asian ethnicity, have genetic variations that prevent them from properly metabolising alcohol.
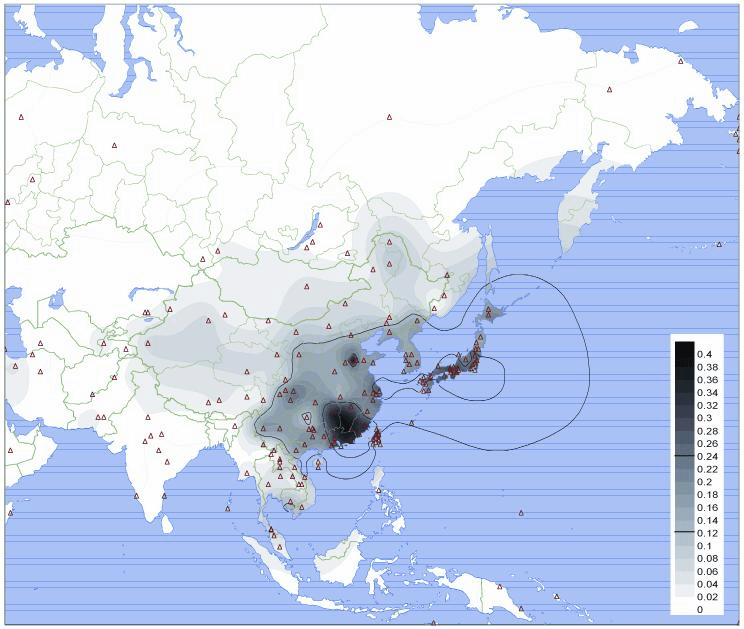
One of the many jobs that the liver performs is the metabolism of alcohol. When we drink it breaks down a lot of the alcohol that is absorbed through the stomach in the so-called first pass effect. The enzyme that catalyses6 the first step in this reaction is called alcohol dehydrogenase and it breaks down alcohol into a very toxic molecule called acetaldehyde. Normally, this molecule is then quickly broken down into acetate by another enzyme in the liver called aldehyde dehydrogenase 2 (ALDH2). People in some Asian populations, and more rarely in other ethnic groupings, have, once again, a SNP in ALDH2 that greatly reduces its ability to break down acetaldehyde. If you have one copy of this variant you are 50% as effective at breaking down acetaldehyde when compared to someone with two functioning variants. If you have two copies you are only 4% as effective.
When people with this version of ALDH2 drink alcohol they quickly get high levels of acetaldehyde in their body which leads to symptoms like flushing, nausea, tachycardia (rapid heartbeat), and headaches. This lowers the risk of alcoholism, because it’s painful to drink, but can lead to increased chances of cancer if people persist in drinking. You may ask why would you drink if it makes you so ill? But, though things have changed somewhat, in many cultures alcohol has a social role. In Japan, for example, where 40-50% of the population have the ALDH2 variant, the workplace custom of the nomikai drinking party has traditionally been a trial for those looking to climb the corporate ladder. Essentially forcing many with the variant to drink, regardless of the deleterious effects.
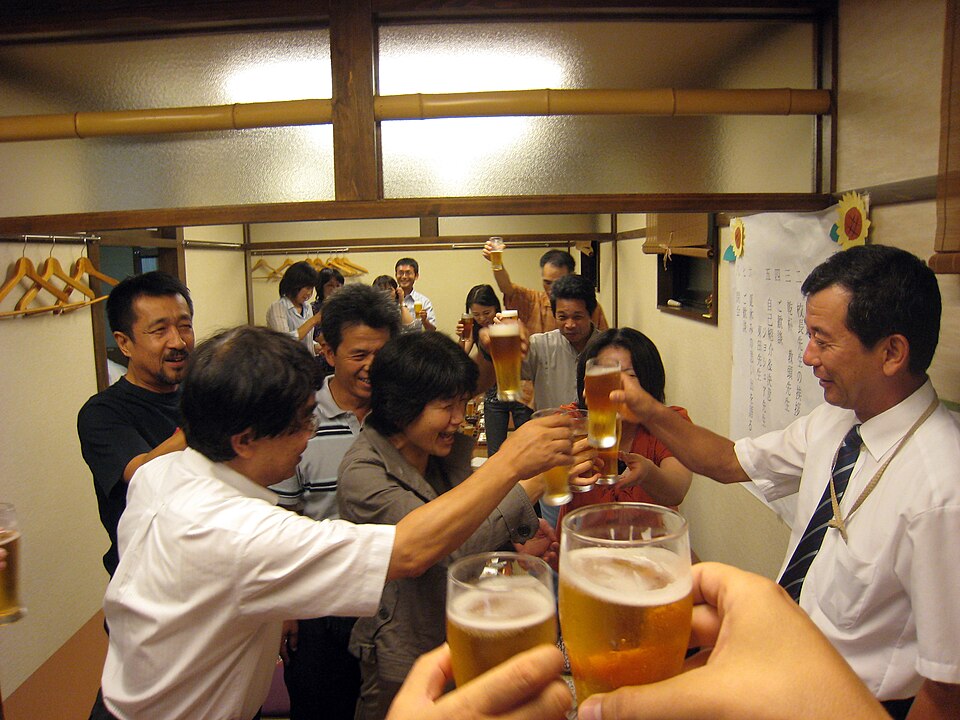

All of the variants we’ve covered here, our flavour preferences, lactose intolerance and ALDH2 are all great examples of how genetic variation affects our relationship with food. They are easy to understand, the biochemistry involved is often relatively simple and they have immediate consequences that are easily linked to their trigger. There are, however, many more variants and some of these are not as easily linked to their consequences. Many gene variants have been associated with obesity, diabetes and cardiovascular disease and these diseases develop over many years. The complex interplay of these gene variants with the food we eat, as well as the other genes in our genome, can make it very difficult to work out exactly how a specific diet, or even specific type of food, will affect a specific individual.
Apart from very broad advice, like eat more vegetables, it is very hard to tell how an individual is going to react to things like paleo, Atkins, keto or the multitude of other dietary fads. We are a long way away from having a good understanding of how genetic variation affects our flavour perception, our food preferences, our metabolism and our health. Anyone who tells you otherwise should be treated with suspicion. You’ll often hear people telling you they’ve “done the research”, normally when they are trying to sell you something. But, when it comes to humans and their genes the only possible outcome of “doing the research” is more confusion. Science often gives us answers but sometimes all it provides is many more questions.
Footnotes
- OK “full” is overdoing it a bit. The human genome has about 3 billion base pairs so 60-70 mistakes probably isn’t “full”. ↩︎
- This is just one of the ways things can go wrong, letters can be deleted, whole segments can be rearranged removed or duplicated and large-scale chromosomal changes can occur. ↩︎
- If you do more reading on genetic diversity in humans you will come across SNPs a lot. ↩︎
- Cilantro if you’re American, Canadian or from Spanish speaking countries. Chinese parsley if you are from some parts of Asia. ↩︎
- I’m simplifying a bit here. There are other genes involved but the end state is that the genetic variations end up being able to detect “soapy” aldehydes at much lower quantities than “normal” human beings. ↩︎
- An enzyme is a protein that makes a reaction proceed quicker than it otherwise would if left to itself. This process of making a reaction go faster is called catalysing the reaction. If you want a primer on this, you can see my post on making fries. It’s about halfway down. ↩︎



Leave a comment